Drug Insight
Reading time required
minutes
Speed reading only takes 9 minutes
👀
4 MicroRNAs: Masters of Gene Regulation
Although researchers demonstrated RNAi functionality using exogenous dsRNAs, RNAi is also induced by the endogenously present single-stranded hairpin microRNAs (miRNAs) in cells. To date, miRNAs are the most abundant species of noncoding RNAs in mammalian cells, and they have fundamental roles in regulating gene expression at the posttranscriptional level. In some cases, RNAi can also confer resistance to virus or other pathogen infection. Notably, the first miRNAs were discovered through forward genetic screens in worms. Indeed, Ambros and colleagues described in 1993 a new mechanism of gene regulation in C. elegans through small non-protein-coding RNA lin-4, which was found to have complementary sequence to the 3′ untranslated region of lin-14 mRNA. In both plants and animals, cloning was the initial strategy of large-scale miRNA discovery. Additionally, several groups developed bioinformatics approaches for miRNA identification. According the miRBase database (release 22.1, http://www.mirbase.org ( http://www.mirbase.org )), there is currently 2656 human mature miRNA sequences that have the capacity to regulate more than two-thirds of protein-coding genes in humans and are involved in every cellular process known so far.
In contrast to siRNAs, most miRNA genes are transcribed in the nucleus by RNA polymerase II, and the primary transcripts known as primary miRNAs (pri-miRNAs) are capped and polyadenylated. Before their export to the cytoplasm for further processing and silencing, nuclear pri-miRNAs must be recognized and cleaved (Fig. 2). These respective tasks are carried out by the proteins Drosha and DGCR8 (known as Pasha in invertebrates), which together constitute the microprocessor complex. As with Dicer, Drosha proteins belong to the RNase III family. The endonucleolytic activity results in the cleavage of the stem structure of pri-miRNAs and the release of the pre-miRNAs, which have a hairpin-like structure. Thereafter, Drosha-generated pre-miRNAs are exported from the nucleus to the cytoplasm by the nuclear export protein Exportin-5, which binds miRNA hairpin duplex along with GTP-bound form of the cofactor Ran, and releases it in the cytoplasm after hydrolysis of GTP. In the cytoplasm, pre-miRNAs are cleaved further by Dicer to generate 22-nt miRNA duplexes containing mismatches and having a two nucleotides 3′ overhang. Subsequently, the miRNA duplex assembles with Ago2 protein to form the effector RISC complex after which the mature miRNA strand is selected (Fig.2). Depending on the homology to mRNAs, miRNA can lead either to mRNA cleavage (complete homology) or to repression of translation (partial homology). In flies and mammals few miRNAs are reported to lead to cleavage of their targets, whereas in plants most miRNAs exhibit complete homology with their corresponding target mRNA genes, resulting in target cleavage. Plant miRNA genes are generally not located within protein-coding genes but comprise their own RNA polymerase II-dependent transcriptional units. This contrasts with animal miRNAs, which sometimes appear to be processed from introns of protein-coding genes. In plants, many predicted miRNA targets encode regulatory proteins, suggesting that plant miRNAs are master regulators. Notably, miRNAs and siRNAs have much in common. Both are 20–24-nucleotides long and processed from longer RNA precursors by Dicer-like ribonucleases, and both are incorporated into ribonucleoprotein silencing complexes in which the small RNAs, through their base-pairing potential, guide target gene repression. However, the fundamental difference between the two small RNA classes is the nature of their precursors; siRNAs are processed from long, double-stranded RNAs, whereas miRNAs are processed from single RNA molecules that include imperfect stem-loop secondary structures as illustrated in Fig.2.
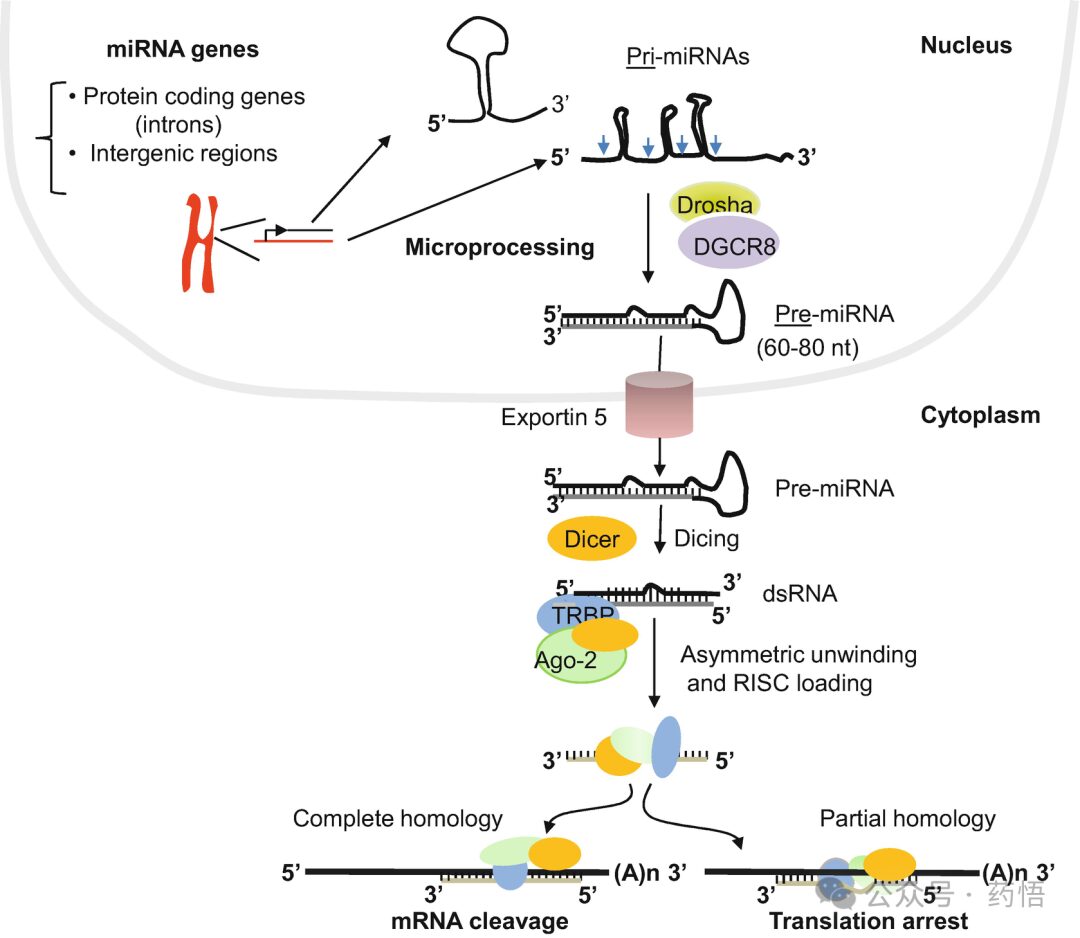
Fig. 2 Schematic representation of miRNA silencing pathway. miRNAs are encoded in the genome and are transcribed to yield primary transcripts (pri-miRNAs), which are processed by Drosha with the aid of its partner DGCR8 protein (Pasha) to generate short precursor miRNAs (pre-miRNAs). These precursor miRNAs are then exported from the nucleus to the cytoplasm, where they are further processed by Dicer to yield miRNA duplexes that were unwound to single-stranded mature miRNA and then loaded into the miRNA-induced silencing complex (miRISC). Unwinding of the complex seems to occur so rapidly after duplex formation due to the Ago2 presence in the complex with Dicer and TRBP. The loaded strand RNA guides the RISC complex to recognize target mRNA sequences. A perfect homology allows Ago-cleavage of mRNA, whereas central mismatches promote repression of mRNA translation.
👀
5 piRNAs: Guardians of the Germline
The Piwi-interacting RNAs (piRNAs) are the most recently discovered class of small non-coding RNAs, and they bind to the PIWI subclade of Argonaute proteins. One well-characterized Piwi/piRNA function is the silencing of retrotransposons, both at the posttranscriptional and epigenetic levels, as well as of other genetic elements in germlines, particularly those during spermatogenesis. Fly piRNAs were initially termed repeat-associated small interfering RNAs (rasiRNAs) because they were found to map to transposons and repetitive elements. Unlike rasiRNAs in flies, the majority of piRNAs in mammals arise from un-annotated regions of the genome devoid of transposons. Currently there are about 23,500 piRNAs identified in the human genome, a number that is quite comparable to that of protein-coding mRNAs (around 20,000), suggesting that piRNAs may have several important roles in gene regulation that remain to be investigated.
Unlike miRNA and siRNA, piRNAs are typically longer, with a length of 26–31, originate from single-stranded RNA precursors, and do not require Dicer for maturation (Fig. 3). Most piRNAs are transcribed by RNA polymerase II as long transcripts which are then exported to the cytoplasm and processed into smaller sequences (mature piRNAs). In D. melanogaster, the nuclear export of piRNA precursors is mediated by UAP-56 that has been previously implicated in mRNA splicing and export. Another factor that has been important in the context of piRNAs biogenesis is the mitochondrial surface protein Zucchini. This protein has endonuclease activity for single-stranded RNA and likely processes the piRNA precursors, possibility at the 5′end. After 5′ processing, the 5′U/piRNA is incorporated into the MID domain of the Piwi protein, followed by 3′end processing, which likely involves an unknown trimmer activity as well as the action of methyl-transferase Pimet. Mature piRNAs are 5′ monophosphated and 2′-O-methyl modified in the 3′ terminal, a unique and specific feature of all piRNAs.
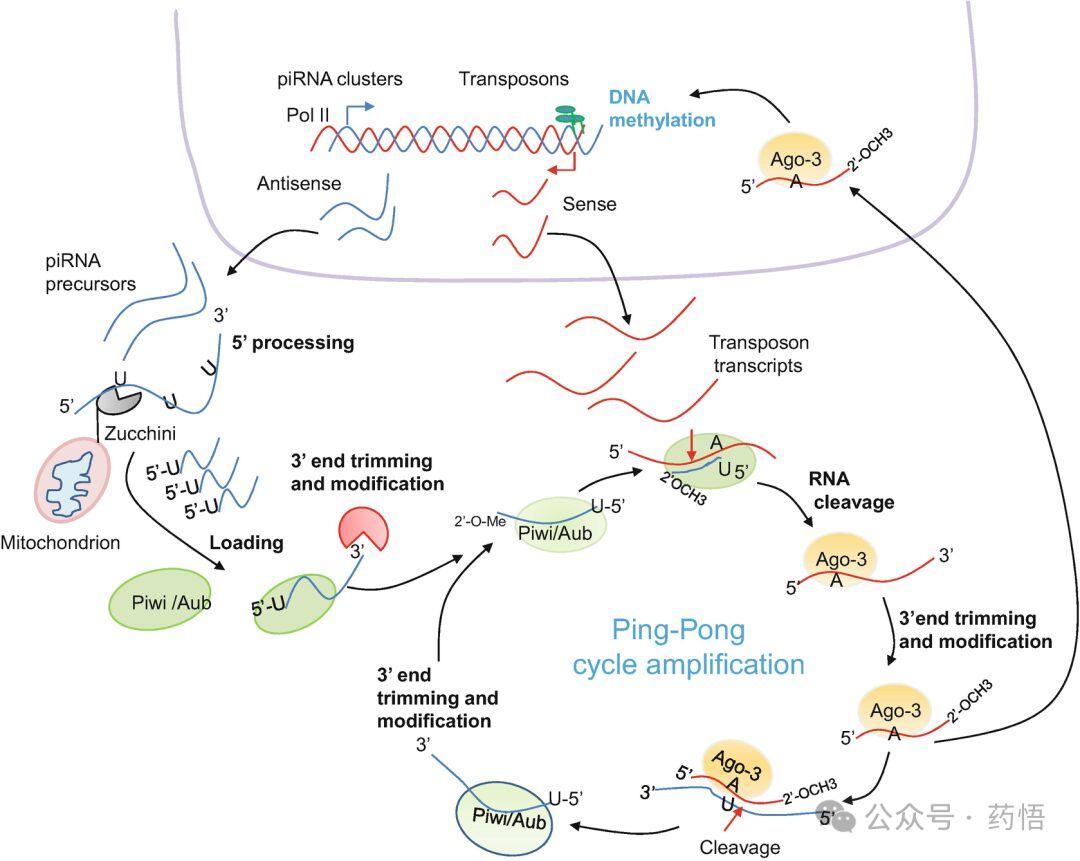
Fig. 3 Mechanisms of piRNA biogenesis in D. melanogaster. The piwi-interacting RNA pathway is involved in defense against transposable elements in the germline. Antisense precursor piRNAs are synthesized from repetitive elements by RNA polymerase II and then exported from the nucleus to nuage granules, where piRNA biogenesis is believed to occur. These piRNA precursors are cleaved by the endonuclease Zucchini, located in the outer membrane of mitochondria, generating the 5′end of the future piRNAs. Then they are trimmed to short fragments, and their 3′ ends are 2′-methylated and then loaded onto aubergine (Aub), which cleaves complementary transposon transcripts to generate sense piRNA fragments. Argonaute 3 (Ago3) uses the sense piRNA fragments to bind and cleave additional complementary antisense transposon transcripts to generate more antisense piRNAs, thus creating a feedback loop known as the ping-pong cycle. This amplification step generates new piRNAs that are identical in sequence to the piRNA that initiated the cycle.
The D. melanogaster genome encodes three Piwi proteins, Piwi, Aubergine (Aub), and Ago3, all of which are required for male and female fertility. As shown in Fig. 3, Piwi and Aub show a strong preference for sequences with a 5′uridine (U) within the antisense to transposons, whereas Ago3 shows enrichment for adenine (A) at position 10 within the sense to transposons, suggesting that Ago3 piRNAs are generated by a new mechanism known as the “ping-pong cycle” of amplification. In this cycle, Aub-bound piRNAs recognize a complementary transposon and induce endonuclease cleavage of the target. Such slicing generates the 5′ end of new sense piRNAs. This cytoplasmic loop, which is also found in zebrafish, links piRNA amplification to posttranscriptional target silencing. Additionally, mature piRNAs in the cytoplasm form a complex with Piwi proteins and migrate back into the nucleus, reaching their target transcripts and mobilizing the silencing machinery to block the transcription of transposable elements via DNA methylation (Fig. 3). Although the role of piRNAs in silencing transposon repeat elements is well established, recent reports have shown that piRNAs can also regulate gene expression and perfect sequence homology between the piRNA and its target mRNA is not required for efficient repression by deadenylation. Apart from the gonads, piRNAs and PIWI proteins may be expressed in various somatic cell types, albeit typically at lower levels than those observed in the germline. Similar to miRNAs, a higher number of piRNAs are expressed in tumors compared to normal tissues.