
OsHDAC1 deacetylates the aldehyde dehydrogenase OsALDH2B1, repressing OsGR3 and decreasing salt tolerance in rice
Research background
Salt stress poses a major threat to rice productivity, significantly impairing growth and yield. Although histone deacetylases (HDACs) are known to mediate plant responses to abiotic stress, the functional role of OsHDAC1 in salt stress tolerance remains poorly understood. In this study, we demonstrate that OsHDAC1 acts as a negative regulator of salt tolerance in rice by deacetylating the metabolic enzyme OsALDH2B1, thereby reducing its stability and downregulating the expression of its downstream target, OsGR3. Genetic evidence supports this regulatory mechanism: knockdown of OsHDAC1 enhanced salt tolerance, whereas overexpression increased sensitivity to salt stress. Further analysis revealed that OsHDAC1 physically interacts with OsALDH2B1 and deacetylates it at lysine residues K311 and K531, promoting its degradation via the ubiquitin-proteasome pathway. Importantly, OsALDH2B1 functions as a transcriptional activator of OsGR3, binding directly to its promoter to induce expression. Since OsGR3 encodes glutathione reductase – a key in ROS detoxification – this regulation is critical for oxidative stress management. Under salt stress conditions, OsHDAC1 expression is downregulated, leading to increased acetylation and stabilization of OsALDH2B1, which in turn upregulates OsGR3 and enhances ROS scavenging. Our findings uncover a novel OsHDAC1-OsALDH2B1-OsGR3 regulatory axis that modulates salt tolerance enzyme in rice.
Research results
1: OsHDAC1 negatively regulates salt tolerance in rice
To assess the role of OsHDAC1 in salt stress response, we compared the performance of OsHDAC1-RNAi (Ri), overexpression (OE), and wild-type Nipponbare (NIP) seedlings under 150 mM NaCl treatment. Following 10 days of salt stress and subsequent recovery, OsHDAC1-Ri plants exhibited superior growth vigor compared to NIP controls, while OsHDAC1-OE plants showed severe wilting symptoms (Fig. 2A). Quantitative analysis revealed the survival rate of OsHDAC1-Ri plants was significantly higher than both NIP and OsHDAC1-OE lines (Fig. 2B-C). Consistent with these observations, OsHDAC1-Ri plants maintained significantly higher relative fresh weight (FW) and dry weight (DW) under salt stress compared to NIP, whereas OsHDAC1-OE plants showed reduced biomass accumulation.
Since salt stress typically induces oxidative damage through reactive oxygen species (ROS) accumulation, we examined oxidative stress markers. Histochemical staining demonstrated stronger DAB (H₂O₂) and NBT (O₂⁻) signals in OsHDAC1-OE plants compared to NIP, while OsHDAC1-Ri lines exhibited stain reduced intensity (Fig. 2D-E). Malondialdehyde (MDA) levels, an indicator of lipid peroxidation, showed similar trends: while all genotypes had comparable baseline levels, salt treatment induced significant MDA accumulation in NIP and OsHDAC1-OE plants but not in OsHDAC1-Ri plants (Fig. 2F). Quantitative H₂O₂ measurements confirmed these findings, with OsHDAC1-OE plants accumulating the highest levels under stress, while OsHDAC1-Ri maintained relatively low concentrations (Fig. 2G).
Collectively, these results demonstrate that OsHDAC1 negatively regulates rice salt tolerance, likely through promoting ROS accumulation and subsequent oxidative damage. The improved stress tolerance in OsHDAC1-Ri plants correlates with reduced oxidative stress markers, suggesting OsHDAC1 functions as a negative regulator in the rice salt stress response pathway.
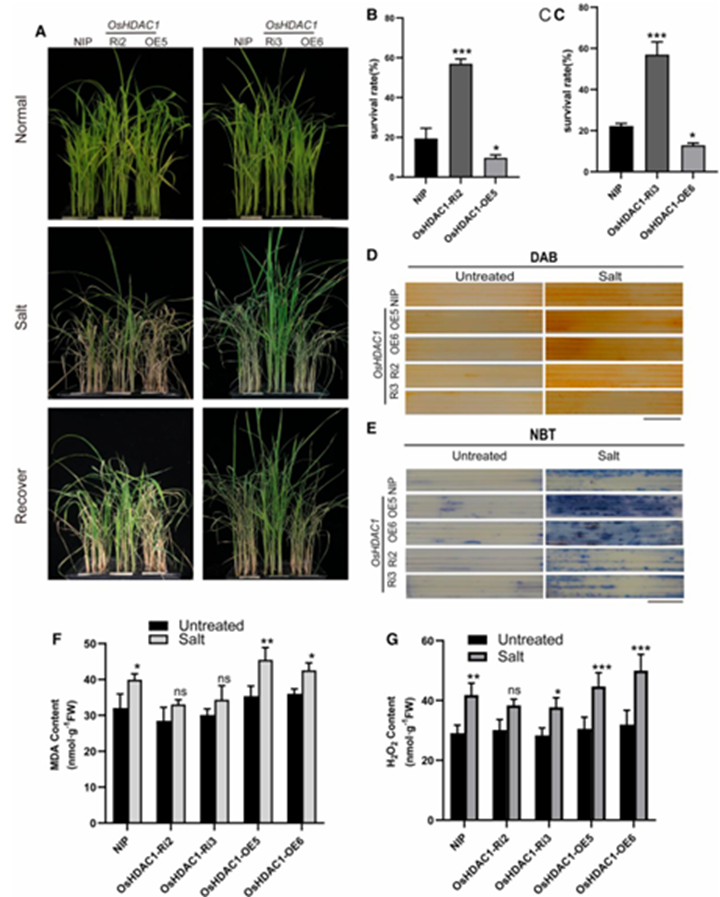
Fig 2
2: OsHDAC1 interacts with OsALDH2B1 and is deacetylated
To investigate the molecular mechanism underlying OsHDAC1-mediated salt stress response, we employed yeast two-hybrid screening and identified acetaldehyde dehydrogenase OsALDH2B1 as a direct interaction partner (Fig. 3A). This physical interaction was further confirmed through multiple independent approaches: (1) luciferase complementation imaging assays demonstrated strong interaction signals (Fig. 3B), (2) bimolecular fluorescence complementation (BiFC) revealed in planta protein-protein interactions (Fig. 3C), and (3) in vitro pull-down assays verified direct binding (Fig. 3D). Importantly, co-immunoprecipitation (Co-IP) experiments in rice protoplasts provided conclusive in vivo evidence for this interaction (Fig. 3E). Collectively, these comprehensive biochemical analyzes establish that OsHDAC1 and OsALDH2B1 interact directly in the nuclear compartment.
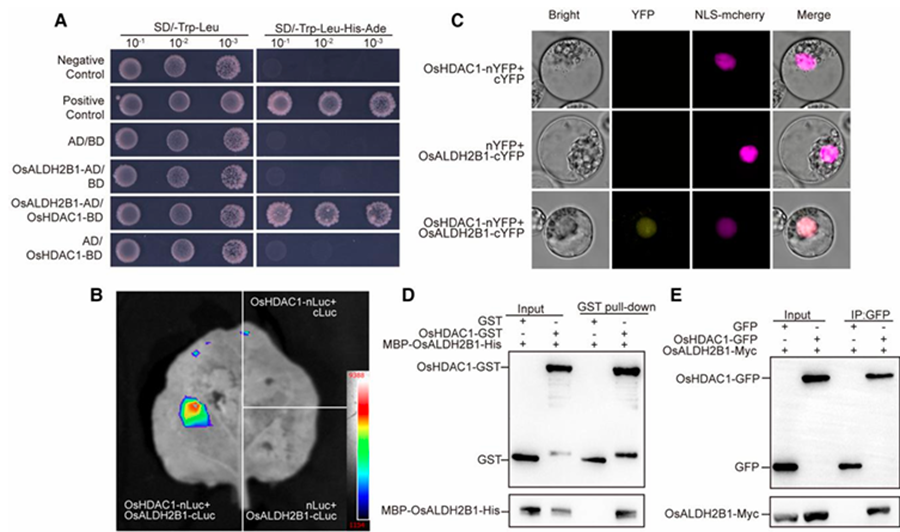
Fig 3
In vitro deacetylation demonstrated that recombinant OsHDAC1-GST protein significantly decreased acetylation levels of OsALDH2B1-His (Fig. 4A). This finding was further validated in vivo using rice protoplasts, where OsHDAC1 similarly reduced OsALDH2B1 acetylation (Fig. 4B). LC-MS/MS analysis identified two specific lysine residues (K311 and K531) as major deacetylation sites on OsALDH2B1 (Fig. 4C). Evolutionary analysis revealed K311 is highly conserved across ALDH family members in both rice and Arabidopsis, while K531 appears unique to OsALDH2B1, suggesting both conserved and protein-specific regulatory mechanisms. To investigate the functional consequences of deacetylation, we examined OsALDH2B1 expression in transgenic plants. Western blot analysis showed reduced OsALDH2B1 protein levels in OsHDAC1-OE plants and Elevated levels in OsHDAC1-Ri lines, while qPCR analysis confirmed mRNA levels remained unchanged (Fig. 4D-E). These results demonstrate that OsHDAC1-mediated deacetylation specifically affects OsALDH2B1 protein stability without altering its transcription, establishing a post-translational regulatory mechanism.
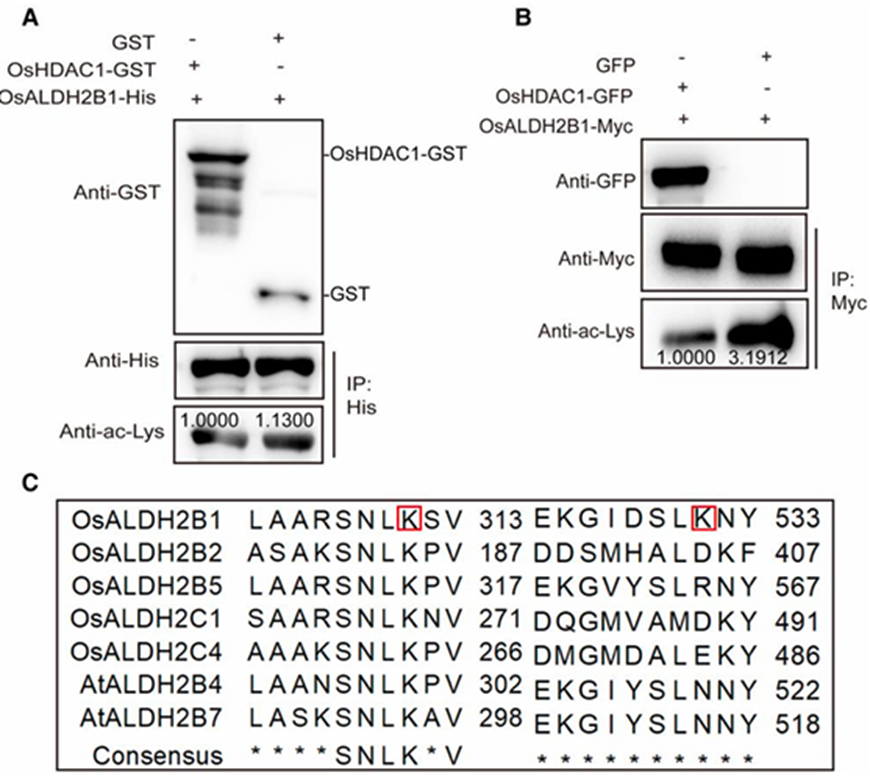
Fig 4
Protein stability assays revealed that crude extracts from OsHDAC1-OE plants significantly enhanced the degradation of recombinant OsALDH2B1-His, whereas extracts from OsHDAC1-Ri plants exhibited a stabilizing effect (Fig. 5A-B). This degradation process was completely abolished by MG132 treatment, confirming proteasome-dependent regulation (Fig. 5A-B). In vivo degradation kinetics further demonstrated accelerated OsALDH2B1 turnover in OsHDAC1-OE lines and prolonged stability in OsHDAC1-Ri plants (Fig. 5C-D). Notably, treatment with trichostatin A (TSA), a potent HDAC inhibitor, substantially delayed OsALDH2B1 degradation, supporting the acetylation-dependent nature of this regulation. To directly investigate the role of specific lysine residues, we generated acetylation-mimetic (K311Q/K531Q, KQ) and deacetylation-mimetic (K311R/K531R, KR) mutants. Stability assays showed the KQ mutant exhibited significantly greater protein stability compared to the KR variant (Fig. 5E-F), providing direct evidence that acetylation status at these sites determines OsALDH2B1 stability.
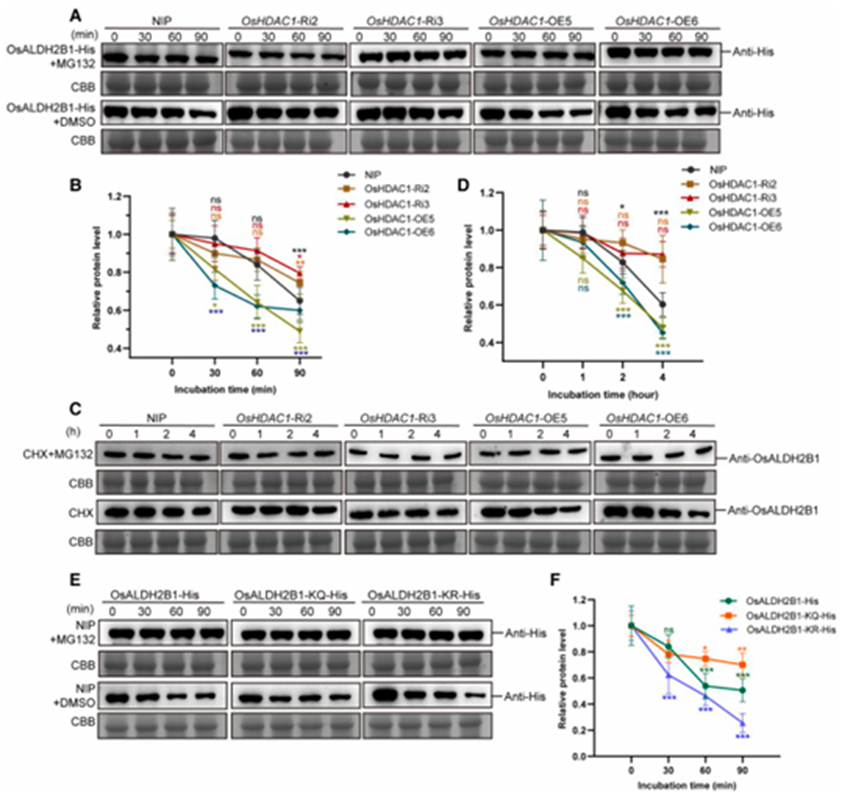
Fig 5
3: OsALDH2B1 positively regulates salt tolerance and activates OsGR3 expression
To investigate the functional role of OsALDH2B1 in salt stress response, we generated and characterized multiple genetic materials: overexpression lines (OsALDH2B1-OE), RNAi knockdown lines (OsALDH2B1-Ri8 and Ri11), and osaldh2b1 knockout mutants. Salt tolerance assays demonstrated striking phenotypic differences – the osaldh2b1 mutant displayed extreme salt sensitivity with rapidly declining survival rates, a phenotype that was partially rescued by treatment with the HDAC inhibitor trichostatin A (TSA) (Fig. 6A-C). Conversely, OsALDH2B1-OE plants exhibited significantly enhanced salt tolerance compared to wild-type Nipponbare (NIP), while both OsALDH2B1-Ri lines showed sensitivity comparable to the knockout mutant.
Consistent with these observations, oxidative stress markers revealed distinct patterns among the genotypes following 6-day exposure to 150 mM NaCl:
Histochemical staining showed substantially stronger DAB (H₂O₂, Fig. 6D) and NBT (O₂⁻, Fig. 6E) signals in OsALDH2B1-Ri lines compared to NIP
· OsALDH2B1-OE plants maintained the lowest oxidative damage markers
· Malondialdehyde (MDA) content, indicative of lipid peroxidation, was significantly elevated in NIP and OsALDH2B1-Ri plants (Fig. 6F)
· While basal H₂O₂ levels were similar across genotypes, salt stress induced dramatic H₂O₂ accumulation in OsALDH2B1-Ri lines but only modest increases in OsALDH2B1-OE plants (Fig. 6G)
These genetic and biochemical analyzes establish OsALDH2B1 as a positive regulator of rice salt tolerance, with its protective effects mediated through mitigation of oxidative stress. The TSA-mediated rescue of the knockout mutant further suggests functional interconnection between acetylation status and OsALDH2B1 activity in stress responses.
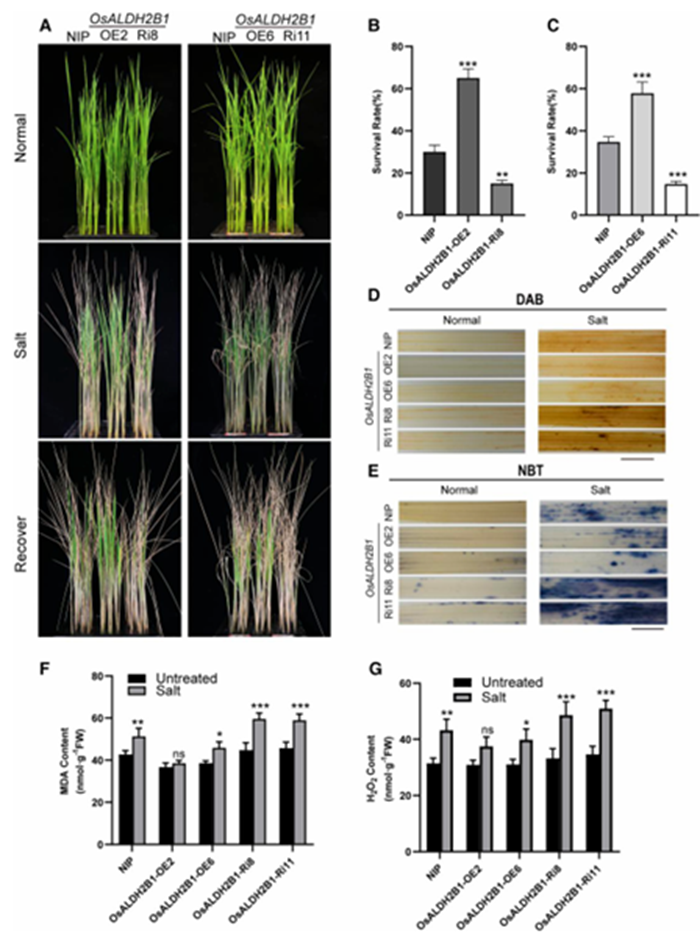
Fig 6
To establish the genetic relationship between OsHDAC1 and OsALDH2B1 in salt regulation tolerance, we generated dual-knockdown lines (OsALDH2B1/OsHDAC1-Ri7 and Rip) in the OsHDAC1-Ri3 background. Strikingly, simultaneous suppression of both genes completely reversed the salt-tolerant phenotype observed in the single OsHDAC1-Ri3 line (Fig. 7). This genetic complementation analysis demonstrates:
1. The salt tolerance conferred by OsHDAC1 knockdown is strictly dependent on OsALDH2B1 function.
2. OsALDH2B1-mediated salt tolerance requires regulation by OsHDAC1.
3. Both genes operate in the same genetic pathway for salt stress response.
These findings provide compelling genetic evidence that OsHDAC1 and OsALDH2B1 function interdependently to regulate rice salt tolerance, with OsHDAC1 acting through post-translational control of OsALDH2B1 stability and activity.
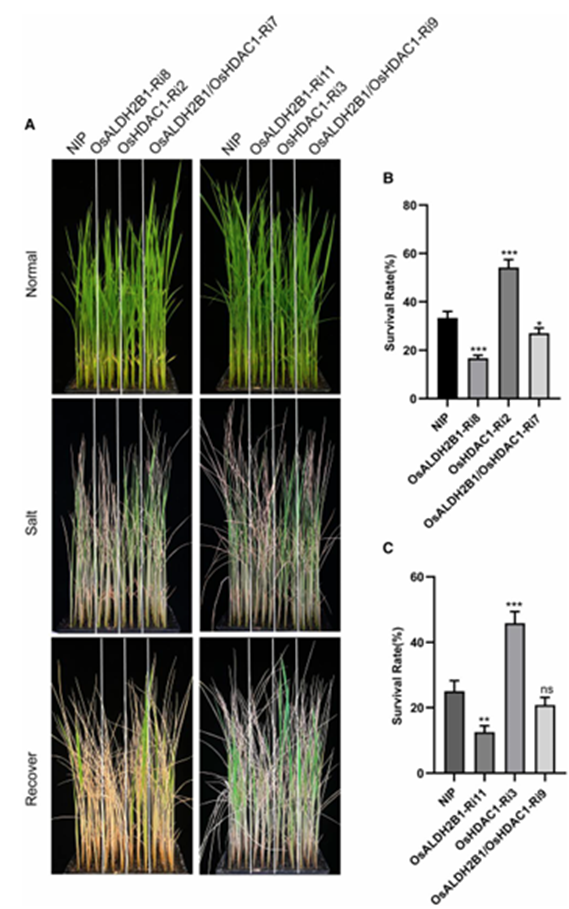
Fig 7
To elucidate the molecular mechanism underlying OsALDH2B1-mediated salt tolerance, we performed comprehensive analysis of its transcriptional regulation of OsGR3, a key glutathione reductase gene. Bioinformatics screening of the 2-kb OsGR3 promoter region identified three potential OsALDH2B1 binding motifs (Fig. 8A). Subsequent electrophoretic mobility shift assays (EMSAs) demonstrated that recombinant OsALDH2B1 protein specifically binds to two distal motifs (ProA and ProB), while showing no affinity for the proximal motif near the transcription start site (Fig. 8B). This specific binding was further confirmed through yeast one-hybrid assays (Fig. 8C).
Functional validation using dual-luciferase reporter assays revealed:
-
OsALDH2B1-Myc strongly activated LUC expression driven by the OsGR3 promoter
-
Co-expression of OsHDAC1-HA significantly suppressed this transcriptional activation (Fig. 8D,E)
Consistent with these findings, RT-qPCR analysis showed:
-
Salt stress induced OsGR3 upregulation in wild-type plants
-
OsGR3 expression was significantly enhanced in OsHDAC1-Ri and OsALDH2B1-OE lines
-
OsHDAC1-OE plants showed no significant change in OsGR3 levels
-
OsALDH2B1-Ri lines exhibited markedly reduced OsGR3 expression
These results establish that:
1. OsALDH2B1 functions as a transcriptional activator of OsGR3 through direct promoter binding
2. OsHDAC1 negatively regulates this activation, likely through deacetylation-mediated control of OsALDH2B1 stability.
3. The OsALDH2B1-OsGR3 regulatory axis represents a crucial component of the rice salt stress response.
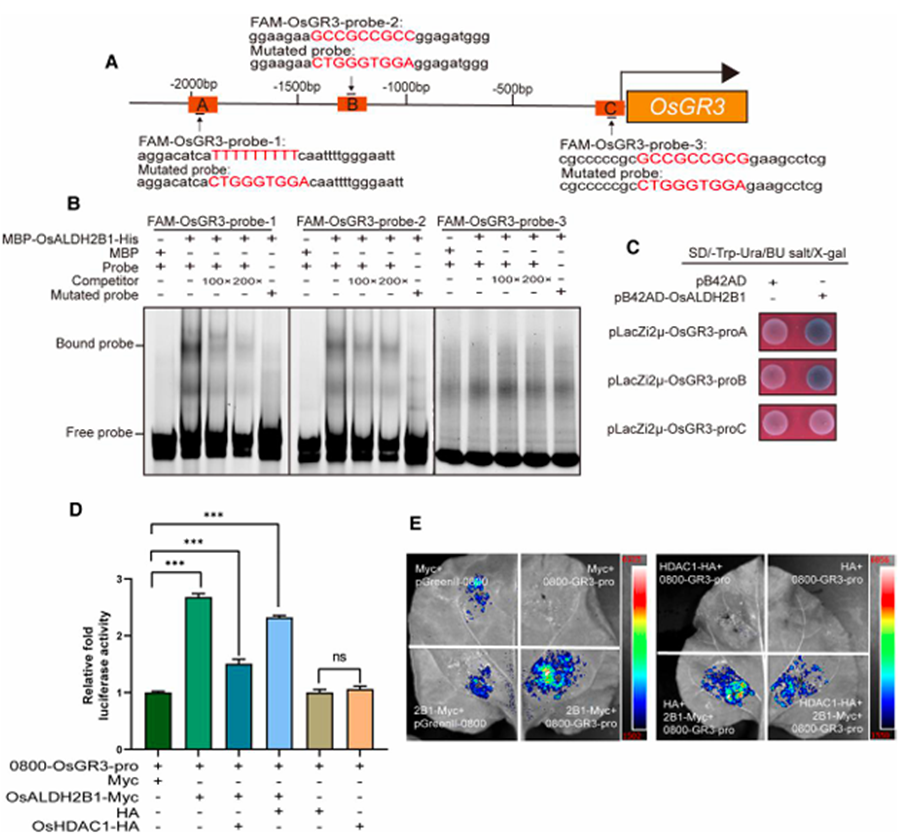
Fig 8
4: Down-regulation of OsHDAC1 expression under salt stress promoted acetylation of OsALDH2B1
To investigate the dynamic regulation of OsALDH2B1 acetylation under salt stress, we first analyzed the temporal changes in OsHDAC1 and OsALDH2B1 protein levels. Western blotting revealed a progressive decline in OsHDAC1 abundance (Fig. 9A), while OsALDH2B1 protein levels increased significantly over time (Fig. 9B). Notably, immunoblotting with anti-acetyl-lysine antibodies demonstrated a corresponding accumulation of acetylated OsALDH2B1 (Fig. 9C), suggesting that salt stress promotes OsALDH2B1 stabilization through enhanced acetylation.
To directly assess OsHDAC1’s role in this process, we compared acetylated OsALDH2B1 levels in wild-type (NIP) and OsHDAC1-Ri plants. Under normal conditions, OsHDAC1-Ri lines already exhibited elevated OsALDH2B1 acetylation compared to NIP. This difference became particularly pronounced after 12 h of salt treatment (Fig. 9D), demonstrating that:
1. OsHDAC1 is the primary deacetylase regulating OsALDH2B1 acetylation status
2. Salt stress actively suppresses OsHDAC1 expression/activity.
3. The resulting hyperacetylation of OsALDH2B1 represents a key adaptive response to salinity
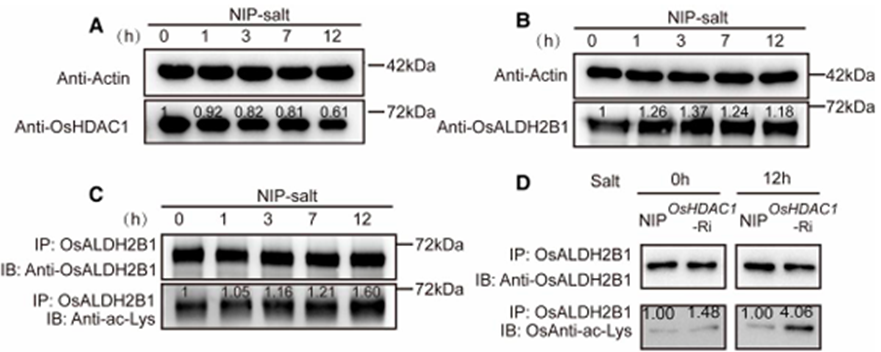
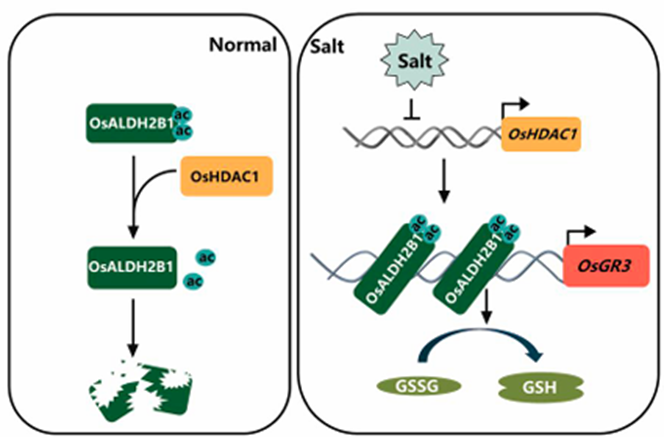
Fig 9
Summary
The findings demonstrate a dynamic, acetylation-dependent regulatory mechanism governing rice salt tolerance. Under normal conditions:
-
OsHDAC1 constitutively interacts with and deacetylates OsALDH2B1 at K311/K531 residues.
-
Deacetylated OsALDH2B1 undergoes ubiquitin-proteasome degradation, maintaining low basal levels
Under salt stress:
1. Transcriptional suppression of OsHDAC1 leads to:
-
Accumulation of acetylated OsALDH2B1
-
Stabilization of OsALDH2B1 protein
2. Acetylated OsALDH2B1 functions as a transcriptional activator:
-
Binds specifically to the OsGR3 promoter (ProA/ProB regions)
-
Upregulates OsGR3 expression, enhances glutathione reductase activity
3. Enhanced glutathione cycling:
-
Increased conversion of GSSG → GSH maintains redox homeostasis
-
Reduced ROS accumulation protects against oxidative damage
This work establishes that salt-induced acetylation switches convert OsALDH2B1 from a degradation-targeted protein to a functional transcription factor, revealing a novel stress adaptation strategy where metabolic enzymes acquire transcriptional regulatory functions through post-translational modification.
edit| Tahreem Li Mu